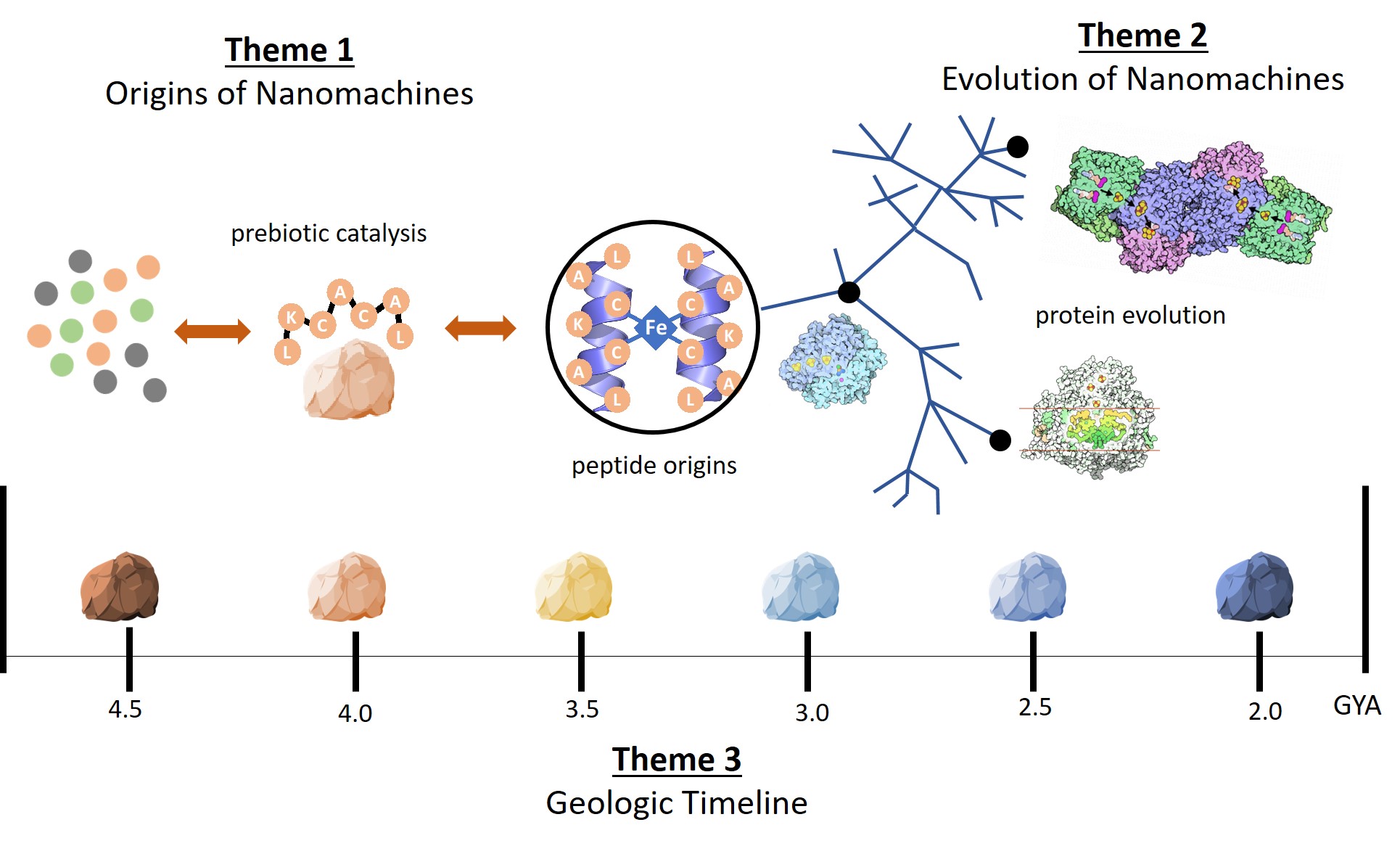
ENIGMA: Evolution of Nanomachines In Geospheres and Microbial Ancestors. This NAI team will explore catalysis of electron transfer reactions by prebiotic peptides to microbial ancestral enzymes to modern nanomachines, integrated over four and a half billion years of Earth’s changing geosphere. Theme 1 focuses on the synthesis and function of the earliest peptides capable of moving electrons on Earth and other planetary bodies. Theme 2 focuses on the evolutionary history of “motifs” in extant protein structures. Theme 3 focuses on how proteins and the geosphere co-evolved through geologic time.
About
Life on Earth is electric. The electronic circuitry is catalyzed by a small subset of proteins that function as sophisticated nanomachines. Currently, very little is known about the origin of these proteins on Earth or their evolution in early microbial life. To fill this knowledge gap, the ENIGMA research team proposes to conduct integrated and coordinated experimental, bioinformatic, and data-driven studies to explore the origin of catalysis, the evolution of protein structures in microbial ancestors, and the co-evolution of proteins and the geosphere. ENIGMA has three integrated research themes focused on understanding the evolution of proteins involved in electron transfer and energy generation:
Synthesis of Nanomachines in the Origin of Life
Theme 1 seeks to understand how complex extant nanomachines that catalyze electron transfer emerged from simple prebiotic chemical processes. Two possible biochemical origins scenarios will be explored: on the early Earth at the beginning of the Archean eon, and on other planets where different amino acid alphabets and chemical constraints might likely be present. Central to both scenarios is the hypothesis that the first functional molecules were small low-complexity metal-binding peptides that were capable of primitive electron transfer and catalysis, and that these functional peptides subsequently evolved into larger proteins. Since no direct physical fossils of these earliest peptides exist, we will instead turn to databases containing thousands of high resolution extant protein structures and powerful computer simulation tools for engineering small peptides from basic physical principles. This will essentially re-invent plausible candidates for early-life peptides.
In Theme 1, terrestrial peptides will be constructed from the twenty natural amino acids, whereas extraterrestrial peptides will incorporate a much broader alphabet, informed by amino acid distributions found in meteorites and prebiotic chemical simulations such as the classic Urey-Miller experiments. Early peptides are proposed to have assembled into circuits capable of conducting electricity where electron transfer occurred primarily through the metals coordinated by peptides. Thus, it would be critical for peptides to preserve structure and metal binding as these sites assume transient oxidized or reduced excited states. In order to assess their functional potential, candidate molecules will be synthesized and tested for metal binding using a number of spectroscopic, scattering and microscopy methods. High resolution structures will be determined by X-ray diffraction and solution NMR methods to ascertain how similar early peptides are to subsets of extant oxidreductases. A series of functional assays that range from electron transfer and catalysis to the ability to act as metabolic surrogates will be used to evaluate the functional capacity of these small peptides. The resulting peptide designs will be used to root protein evolutionary trees and to inform powerful machine learning tools that will link information gathered on early peptides, extant proteins and Earth’s geochemical cycles across all three Themes.
Increasing Complexity of Nanomachines in Microbial Ancestors
Theme 2 will examine the emergent complexity of metalloproteins in microbial ancestors. We will develop new computational methods for linking protein structure to the evolution of function, particularly to redox functionality of metal binding proteins. Methods for pairwise functional comparisons of metal-binding folds will be systematically constructed used data from the Protein Data Bank (PDB). We will then use available evolutionary tree-building techniques, augmented with newly developed approaches, to evaluate relative time of appearance of all folds in our set.
Internal tree nodes, the imputed potential ancestral folds of extant proteins, will be synthesized in the lab to verify the tree topology. The composition of ancestral folds in terms of Theme 1 motifs, as well as their function and structure will also be assessed. Finally, we will design computational methods for mapping mineral/metal/trace-element availability to our evolutionary tree. In this manner we will be able to identify the oldest metal-binding protein structures and to time the early evolution of metal binding folds. The machine-learning component of this final task will also allow us to identify trace-element/metal signatures indicative of the various stages of development of life.
Co-Evolution of Nanomachines and the Geosphere
Theme 3 will explore the co-evolution of minerals and proteins through geologic time. We propose to carry out data-driven abductive studies of mineral evolution that document the changes in Earth’s mineral diversity and distribution through deep-time, focusing on mineral interactions with the biosphere. Mineralogical data (e.g., mineral composition, solubility, and age) will be incorporated into geochemical models to constrain the concentration of metals over geologic time and metal bioavailability during Archean and Proterozoic eons.
We will then use the availability of redox-active transition metals to date specific positions in the protein evolution tree constructed in Theme 2. Employing large meta-omics and geochemical datasets of modern environments, we will test if metal concentrations are reliable markers for the occurrence of protein cofactors, and thus the validity of using metal bioavailability to constrain a protein evolution timeline. Finally, we will explore if the co-evolution of minerals and proteins resulted in specific suites of minerals that may actually be fossil “geobiosignatures” of life. We hypothesize that mineral diversification on Earth was a consequence, direct or indirect, of the biosphere.
Using a “Large Number of Rare Events” statistical approach, we will compare Earth’s mineral distribution with that of Mars, the Moon, and meteorites. This work will allow us to distinguish key differences in mineral diversity and distribution on terrestrial planets and moons, and reveal if minerals require life to proliferate.
The proposed team is dedicated to understanding how nature formed catalysts that serve as the pervasive nanomachines of life on Earth, and how analogous processes may have evolved on other planetary bodies within or beyond our solar system. The results of our research will advance our knowledge of how biochemistry emerged from geochemistry—specifically, the enzymatic functions of metal-bearing proteins emerged approximately 4 billion years ago from a geochemical and mineralogical milieu that in some respects mimicked the emergent biochemistry. This concept is of significance to the astrobiology context of all NASA missions, as we attempt to characterize critical features of near-surface environments in which life might emerge. Ultimately, our goal is for the proposed effort to inform future NASA missions about detection of life on planetary bodies in habitable zones. Our effort provides a unique window to potential planetary-scale chemical characteristics that might arise from abiotic chemistry, which must be understood if we are to recognize unique biosignatures on other worlds.

